UVAS provides with these commands for simulations:
Apply_Strain_Profile. [TO BE DEVELOPED]
Load_To_Continue. [TO BE DEVELOPED]
Save_3D_Config. [TO BE DEVELOPED]
Save_To_Continue. [TO BE DEVELOPED]
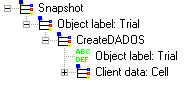
It should be the first command for every simulation. With this command, a silicon simulation box is created as simulation domain.
Parameters:
Object label and Client data: It is not necessary to be worried about it as these words are only used by internal code to work. Only give a name for them.
Parameters file: It is the DDP file where the simulation parameters are defined.
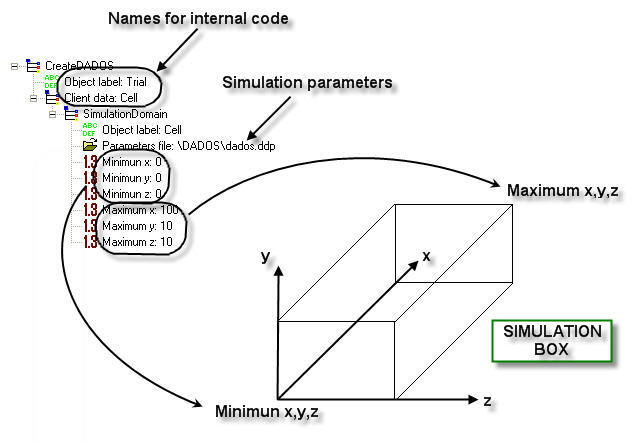
Minimum x, Minimum y, Minimum z, Maximum x, Maximum y, Maximum z: The simulation box used by DADOS to work is simply defined by two 3D points (see figure below), indicating the two opposite sides of the box (distances must be expressed in NANOMETERS). Obviously, the larger the box is, the more accurate and statistically significant the simulation will be, although a longer time will be needed to simulate. DADOS uses as internal minimun x the value zero, this means that if you use a different value for Minimum x, a part of the simulation box will not be shown in the results.
This command will be shown in every command as it is the basic one. It can be checked that when this command has been executed, a tick appears not only in the main command but in all the others.
Click here if you want to learn more about box handling.
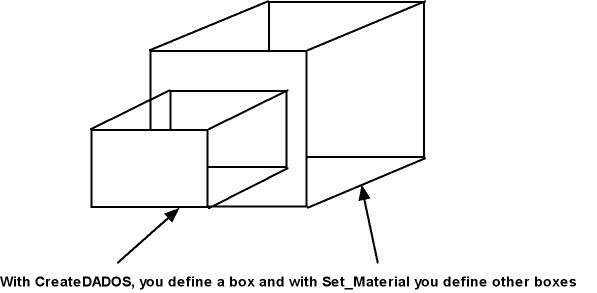
With the command Set_Material, UVAS can create different regions for simulations:
When executed (verbosity level 2), the following is shown in the messages window. As we can see, UVAS shows full information about boxes and amorphization parameters.

The annealing process is simulated by changing the temperature.
Parameters:
Initial Temperature and Final Temperature: UVAS makes a linear interpolation (short steps), as it works with discrete parameters. These temperature must be expressed in DEGREES CENTIGRADE.
Time s: The time between initial and final time must be expressed in SECONDS.
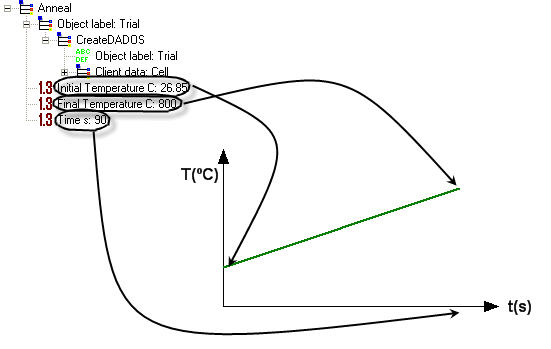
If a constant temperature for annealing is required, please set two annealing processes: the first one will last a little time (e.g. 0.5 s for each 100 °C), from initial temperature up to the final one, and in the second anneal maintain equal both temperatures for the required time:

This technique is not compulsory, a discontinuity in temperature is allowed in DADOS but it is highly recommended setting slopes as shown.
This command will be used to create a simulation box with a variable in-plane lattice parameter depending on the depth (x).

DADOS uses the MARLOWE code, a program based on the binary collisions approach (BCA) in order to generate cascades to implant in silicon (Robinson et al. 1974). After the cascades have been generated, DADOS stores them in an IMPLANT file, which is saved in the ImplantFiles directory. This way, in the next simulation, it is not necessary to re-calculate the cascades. However, if the simulation box is modified, IMPLANT file will be modified as well. It is very important that only one similar implant in the same directory can be simulated at the same time, in order not to overwrite the file.
When executed (verbosity level 2), the following is shown in the messages window. In the example, as it didn't exist the IMPLANT file, UVAS creates it:
![]()
Parameters:
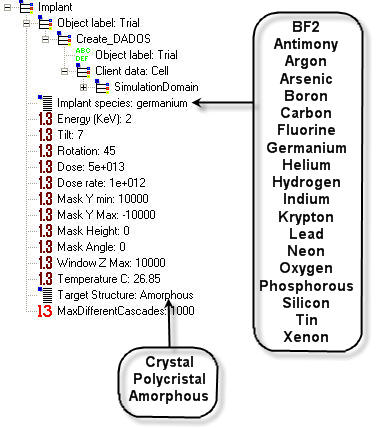
Implant species: A species to implant can be selected here from a list. Not all the species are implemented (see figure below). For those who are not, DADOS only creates the damage. If you want to do the same with other implemented species, please set a negative energy.
Energy: It is the implant energy, expressed in KILOELECTRONVOLTS.
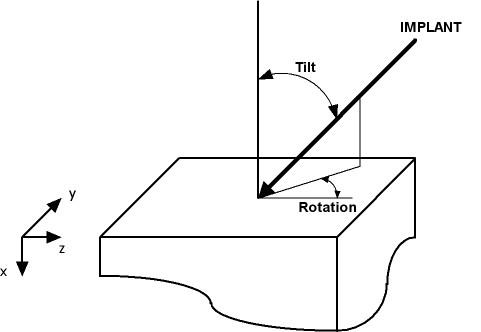
Tilt: See figure below. When tilt is zero, the implant direction is <100>.
Rotation: See figure below.
Dose: Total concentration to implant, expressed in IONS PER SQUARE CENTIMETER.
Dose rate: Implant rate, expressed in IONS PER SQUARE CENTIMETER AND PER SECOND.
Mask Y min: See figure below.
Mask Y Max: See figure below.
Mask Height: See figure below.
Mask Angle: See figure below.
Window Z Max: See figure below.
Temperature: Implant temperature expressed in DEGREES CENTIGRADE. This is an important parameter as implant results (damage) depends on temperature.
Target Structure: A target structure can be selected here from a list.
MaxDifferentCascades: MARLOWE generates the number of cascades indicated by this parameter. DADOS repeats them in different positions randomly chosen. It should be big enough to avoid statistic errors but not huge.
It is obvious that implant time can be obtained as dose/dose_rate, as it is a linear process.
DADOS implements two types of conditions for cascades: periodic and mirror.
Tilt and Rotation definition:
Parameters for the mask:
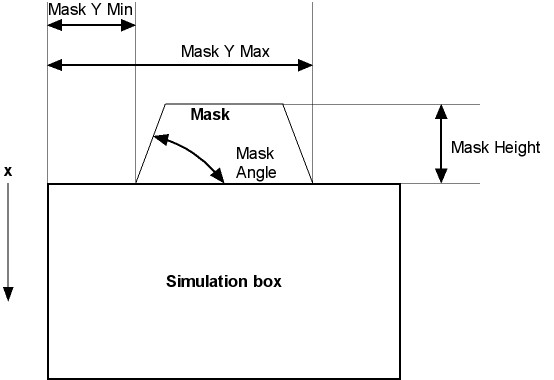
If you want to eliminate the mask, please set Mask Y Min higher than Mask Y Max.
This command should be used at any moment after loading a profile file with Read_Profile. It implants all the profiles opened so far simultaneously.
Parameters:
Time: Time used to implant the profile. This parameter must be expressed in SECONDS. Time can be set to zero, but at high temperatures, if the particles are mobile, a great amount of processes are happening that can affect the computer stability; therefore, it is highly recommended using a non-zero value if the particles are mobile.
Temperature: Implant temperature expressed in DEGREES CENTIGRADE. This is an important parameter as implant results (damage) depends on temperature.
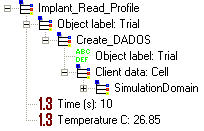
This command is useful for:
A DA2IN file can be inserted here to be plotted as external data for visualization purposes. Please see data file format for more information.
This command must be used with a RESTART file, generated with Save_To_Continue. It is used to continue with a simulation.
When this command is reached during the simulation, UVAS opens the graphics structure defined in the indicated GPH file. Its function is similar to the one provided by Main Menu. Click here to know more about graphics.
This command is useful for creating profiles in silicon. At any moment after this command, Implant_Read_Profile should be called in order to implant it. As shown in the figure, it is necessary to provide a file with the information of the implant as symply as giving two columns: one for depth and the other one for concentration (please see data file format for more information). The particle type is given by the file extension:
| .int | Interstitials |
| .dam | Damage |
| .vac | Vacancies |
| .ars | Arsenic |
| .bor | Boron |
| .car | Carbon |
| .flu | Flourine |
| .hyd | Hydrogen |
| .ind | Indium |
| .oxy | Oxygen |
| .pho | Phosphorus |
| .ant | Antimony |
Once a script has been saved, it can be opened using this command in order to continue with a new simulation after another one have been finished. This way, several simulations can be chained to be executed (at different times, not simultaneously).
Commonly, this command is preceded by a save command, such as Save_Output_Plots or Save_Simulation.
This command saves the 3D particle configuration data to a 3D file. The saved configuration is the one at the moment of this command has been called.
WARNING: In general, it is not recommended to set this command during or just after the implant process, as a lot of memory would be required due to the fact that ion implementation generates a great amount of interstitials and vacancies, anc the required memory to store their coordinates would be very high.
This command is used by UVAS to store the output plots into a GIF file (one per plot). Its function is similar to the one provided by Main Menu. This command uses the program GNUPLOT, located in BIN directory. The name of the file will be the name of the simulation followed by a dot and followed by the name of the graph. For instance, if the name of the simulation is "Amorph_Threshold" and the name of the graphic is "Amorph vs Time", the GIF file will be Amorph_Threshold.Amorph vs Time.gif. Click here to know more about graphics.
This command does not save the entire simulation (for doing this, please use Save_Simulation). However, it can be useful for well-known simulations, in order to save memory.
When UVAS reads this command, the simulation with its current results are saved into an UVA file. Its function is similar to the one provided by Main Menu. The information stored is the present at the moment UVAS reads this command. After a simulation has been saved as an UVA file, it cannot be run any more. For running it again, please use Save_To_Continue command.
With this command, the entire simulation is saved, including results. Notice that a great amount of memory might be required, depending on the output options (number of snapshots, 3D structures, etc.).
Sometimes, simulations are long, specially if simulation box is large. In this case, it can be interesting to pause the simulation after a step. For doing this, just set Save_To_Continue command when pause is required and UVAS will save all the information in the file with RESTART extension. For restarting the simulation, please use Load_To_Continue command with the RESTART file saved before.
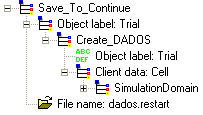
Compare this command with the "Save" function.
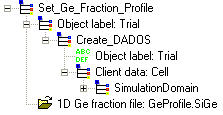
Use this command to set the germanium fraction profile along "x" axis. Information about it must be included in the SIGE file. Click here to know the structure.
It is used to properly define the interface in silicon. If this command is not used, DADOS assumes that there is neither surface nor interface; therefore, there is no simulation of generation or recombination there. Remember that point defects are generated in the surface under equilibrium conditions, thus, the use of this command is highly recommended in most simulations. Generation and recombination in the volume are not implemented in DADOS, as their frequency is extremely low, in order to save memory and computerization resources. Silicon oxide or Ambient are the most common materials employed but other ones can be chosen from those shown below.
Parameters:
Initial x, Initial y, Initial z: Initial coordinates for the new material, expressed in NANOMETERS. Click here to know more about domain definitions.
Final x, Final y, Final z: Final coordinates for the new material, expressed in NANOMETERS. Click here to know more about domain definitions.
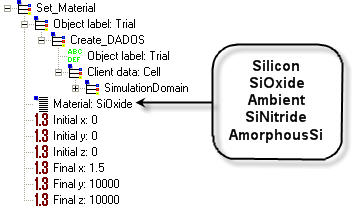
As DADOS works with boxes, and the number of boxes for materials must be integers, it is not guaranteed that the final material size is going to be the specified one but DADOS will do its best. For example, if the box size is X = 100 nm, Y = 10 nm, Z = 10 nm, with 19 bits, the x size of small boxes is 0.78125 nm, which means that the size of material in x axis will be of two boxes, i.e., 1.5625 nm, and not 1.5 nm (the specified one).
This is an important command as UVAS doesn't save every result in order not to waist a lot of memory. It must be set BEFORE the commands which provide results. Here, you will indicate what you want to save. Please click here in order to know more about UVAS data structure.
Parameters:
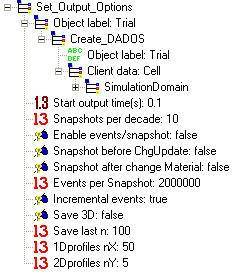
Start output time (s): It is the time when the first snapshot is taken. It has the same function as InitOutputTime in the parameters file, although the value here is the one UVAS will take for the simulation.
Snapshots per decade: Indicates the number of snapshots saved per time decade. If this value is set to zero, UVAS will save a snapshot only when the material changes (if checked this option), or when the charge is updated (if checked this option). Notice that, this way, the snapshot will reflect the worst correlation between majority carriers and dopants. This is useful only for testing the approximation (not valid for physical outputs).
Enable events/snapshot: When selected, UVAS saves a snapshot when a number of events specified in Events per Snapshot happen.
Snapshot before ChgUpdate: When selected, UVAS saves a snapshot just before the charge is updated.
Snapshot after change Material: When selected, UVAS saves a snapshot when the material changes: amorphous to crystalline or crystalline to amorphous.
Incremental events: When selected, UVAS shows the hops and supersaturation averaged over the last snapshot interval. If not, UVAS will average this magnitudes over the whole simulation time.
Save 3D: When selected, UVAS saves 3D structures, useful for plotting atomic configurations. WARNING: In general, it is not recommended to set this command during or just after the implant process, as a lot of memory would be required due to the fact that ion implementation generates a great amount of interstitials and vacancies, anc the required memory to store their coordinates would be very high.
Save last n: Please indicate here the number of snapshots UVAS will save. If more than "n" are generated, UVAS will discard the first ones.
1Dprofiles nX: Number of points in 1D profiles for abscissas. It is not recommended that this parameter is too high in order to save memory.
2Dprofiles nY: Number of points in 2D profiles for ordinates. The number of points in abscissas axis is taken from 1Dprofiles nX. It is not recommended that this parameter is too high in order to save memory.
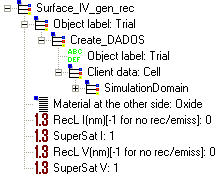
After defining another material by means of Set_Material, it is necessary to define the interface characteristics. This command is also interesting to simulate oxidation and nitridation processes.
Parameters:
RecL I: Recombination length for interstitials. If this value is zero, it means that surface is a perfect sink for interstitials. If this value is set to "-1", the surface will neither recombine nor emit interstitials. This parameter must be expressed in NANOMETERS. DADOS uses a value to initialize, specified in the DDP file, RecLnm.
SuperSat I: It is used so that the interstitials concentration at the interface is different to the equilibrium one. It is useful to simulate oxidation or nitridation processes. DADOS uses a value to initialize, specified in the DDP file, SuperSat.
RecL V: Recombination length for vacancies. If this value is zero, it means that surface is a perfect sink for vacancies. If this value is set to "-1", the surface will neither recombine nor emit vacancies. This parameter must be expressed in NANOMETERS. DADOS uses a value to initialize, specified in the DDP file, RecLnm.
SuperSat V: It is used so that the vacancies concentration at the interface is different to the equilibrium one. It is useful to simulate oxidation or nitridation processes. DADOS uses a value to initialize, specified in the DDP file, SuperSat.
When this command is found in a simulation, UVAS takes a snapshot of current configuration and is saved in the final data structure.